Structure-based protein–ligand interaction fingerprints for binding affinity prediction - ScienceDirect
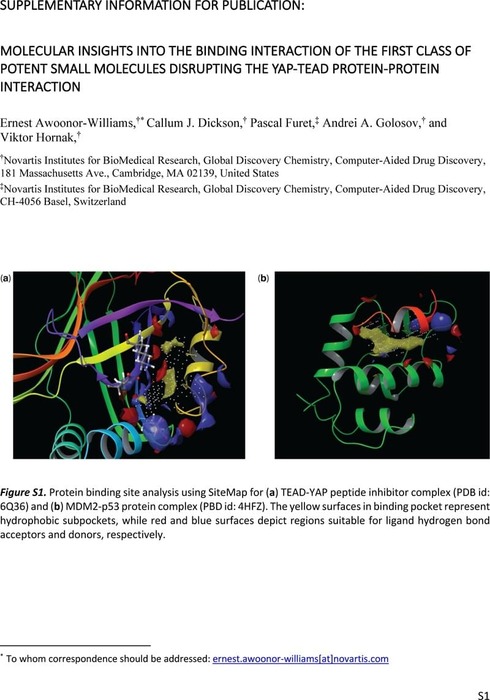
Molecular Insights into the Binding Interaction of the First Class of Potent Small Molecules Disrupting the YAP-TEAD Protein-Protein Interaction | Theoretical and Computational Chemistry | ChemRxiv | Cambridge Open Engage

Three dimensional view of the binding interaction of 3d with active... | Download Scientific Diagram

Binding interactions between compound 5 and the COX-2 binding site. (A)... | Download Scientific Diagram

Intriguing role of water in protein-ligand binding studied by neutron crystallography on trypsin complexes | Nature Communications

Polypharmacology rescored: Protein–ligand interaction profiles for remote binding site similarity assessment - ScienceDirect
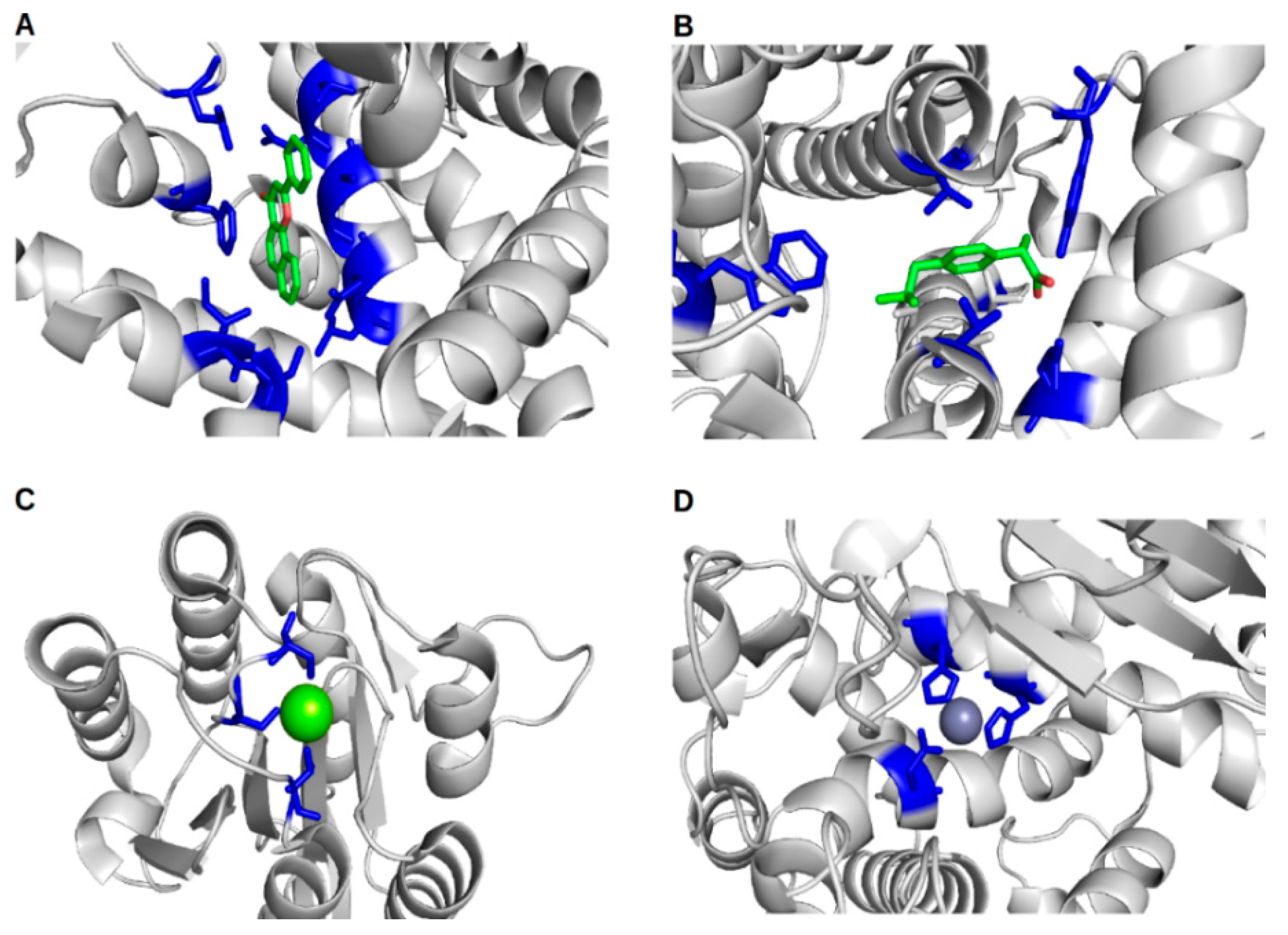
IJMS | Free Full-Text | Proteins and Their Interacting Partners: An Introduction to Protein–Ligand Binding Site Prediction Methods
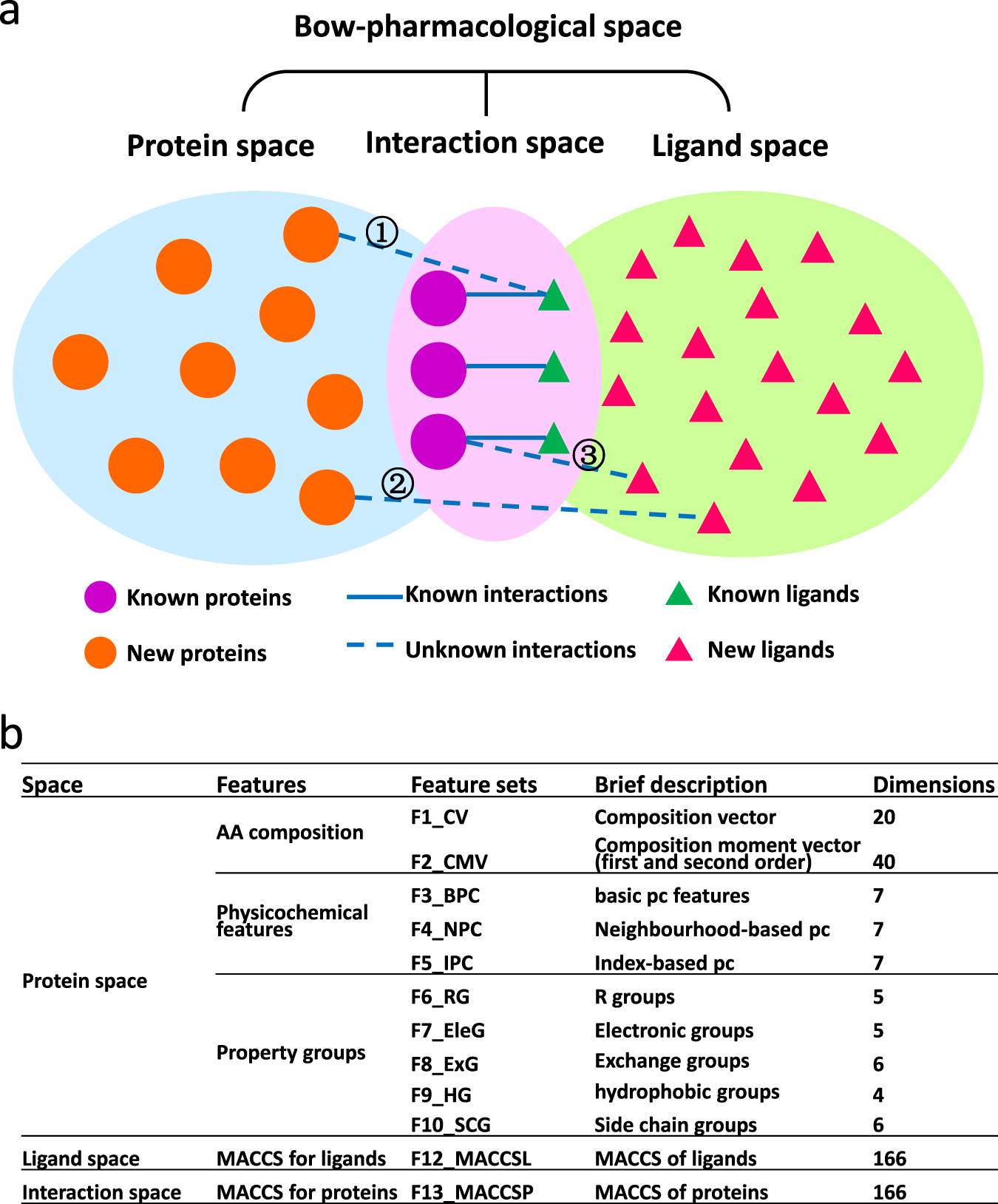
Predicting protein-ligand interactions based on bow-pharmacological space and Bayesian additive regression trees | Scientific Reports

On the binding affinity of macromolecular interactions: daring to ask why proteins interact | Journal of The Royal Society Interface

Geometric Interaction Graph Neural Network for Predicting Protein–Ligand Binding Affinities from 3D Structures (GIGN) | The Journal of Physical Chemistry Letters

Predicting the binding affinities of compound–protein interactions by random forest using network topology features - Analytical Methods (RSC Publishing)

Binding-interaction analysis of (a) QGRG; (b) ashwagandhanolide; (c)... | Download Scientific Diagram
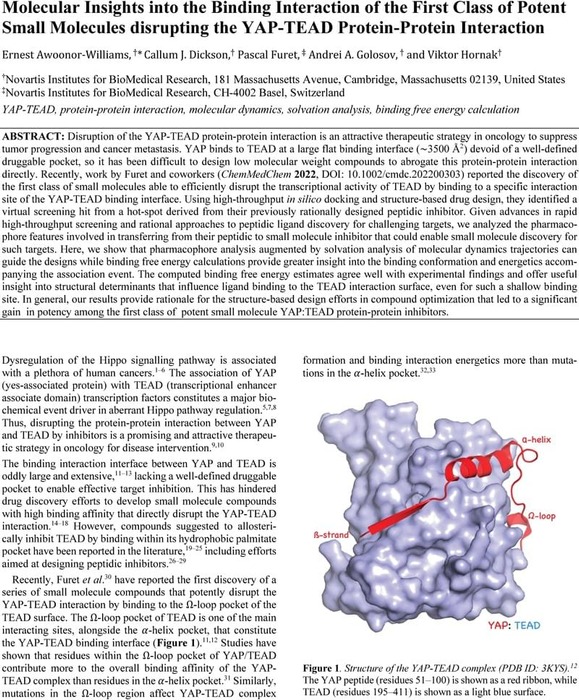
Molecular Insights into the Binding Interaction of the First Class of Potent Small Molecules Disrupting the YAP-TEAD Protein-Protein Interaction | Theoretical and Computational Chemistry | ChemRxiv | Cambridge Open Engage
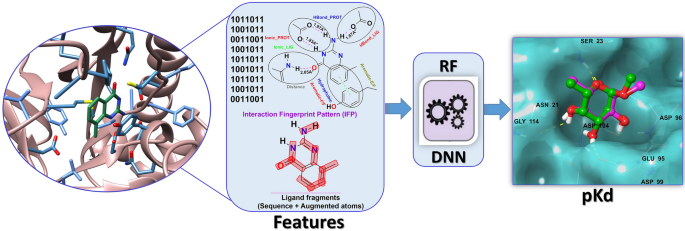
SMPLIP-Score: predicting ligand binding affinity from simple and interpretable on-the-fly interaction fingerprint pattern descriptors | Journal of Cheminformatics | Full Text

Evaluating the binding efficiency of pheromone binding protein with its natural ligand using molecular docking and fluorescence analysis | Scientific Reports

The key binding interactions of compounds 5, 8 and PF-04937139 in the... | Download Scientific Diagram