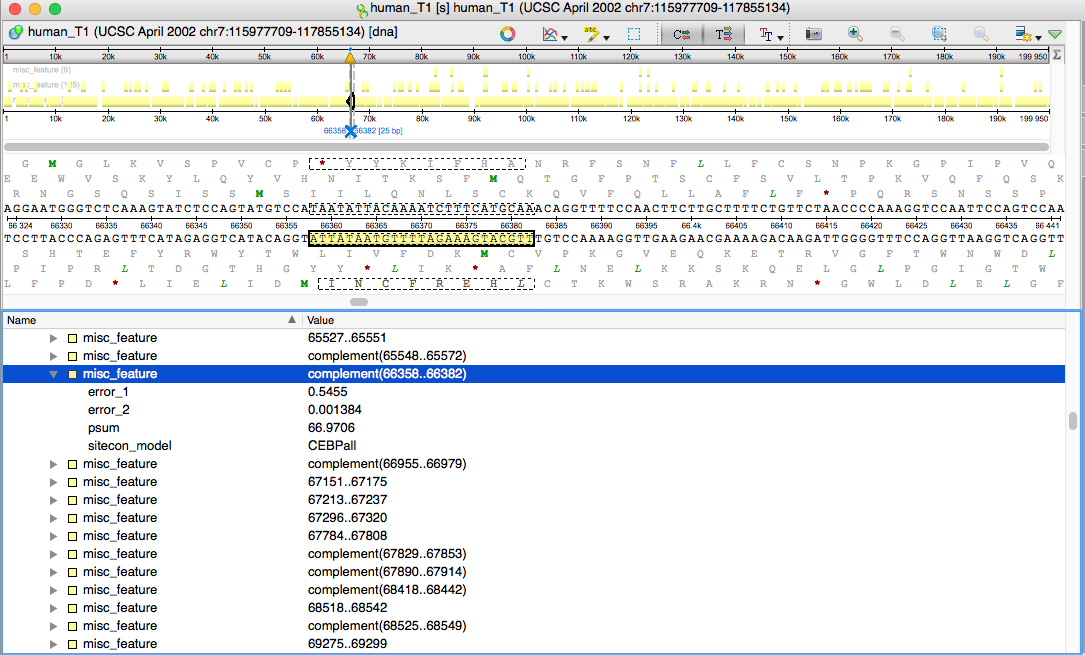
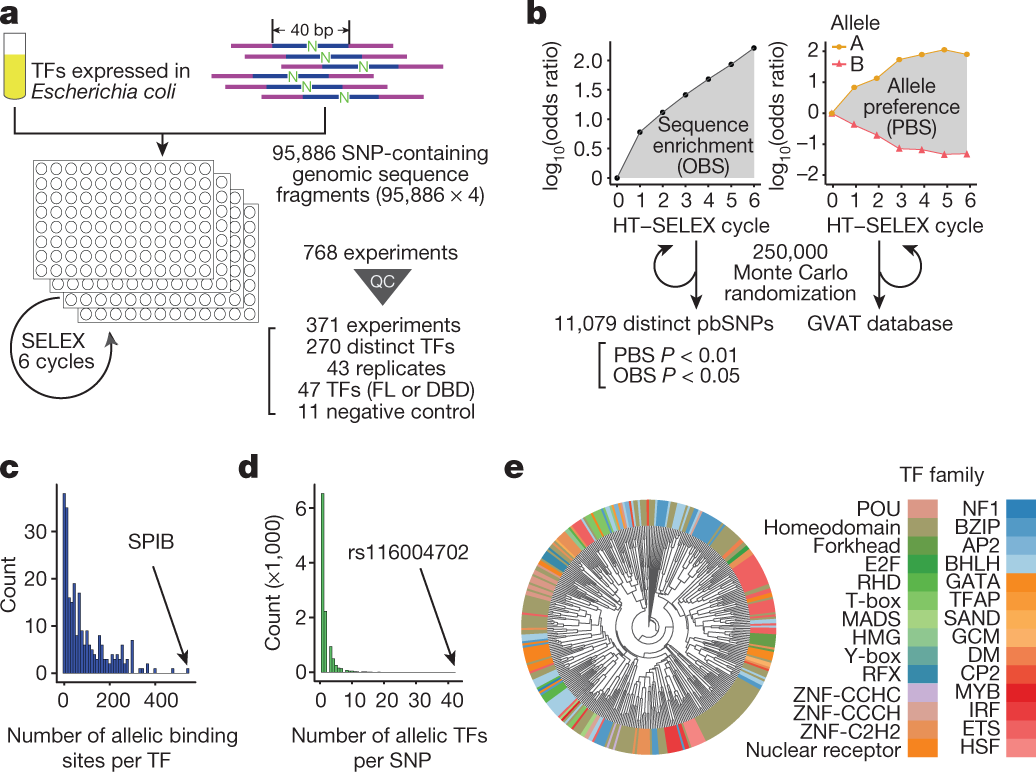
How To Use the Conserved Domain Database (CDD): identify amino acids involved in binding or catalysis
![PDF] sc-PDB: an Annotated Database of Druggable Binding Sites from the Protein Data Bank | Semantic Scholar PDF] sc-PDB: an Annotated Database of Druggable Binding Sites from the Protein Data Bank | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/7b4533a875fde931b28bf1906eda984350587e60/2-Figure1-1.png)
PDF] sc-PDB: an Annotated Database of Druggable Binding Sites from the Protein Data Bank | Semantic Scholar

Automatic generation of bioinformatics tools for predicting protein-ligand binding sites. - Abstract - Europe PMC

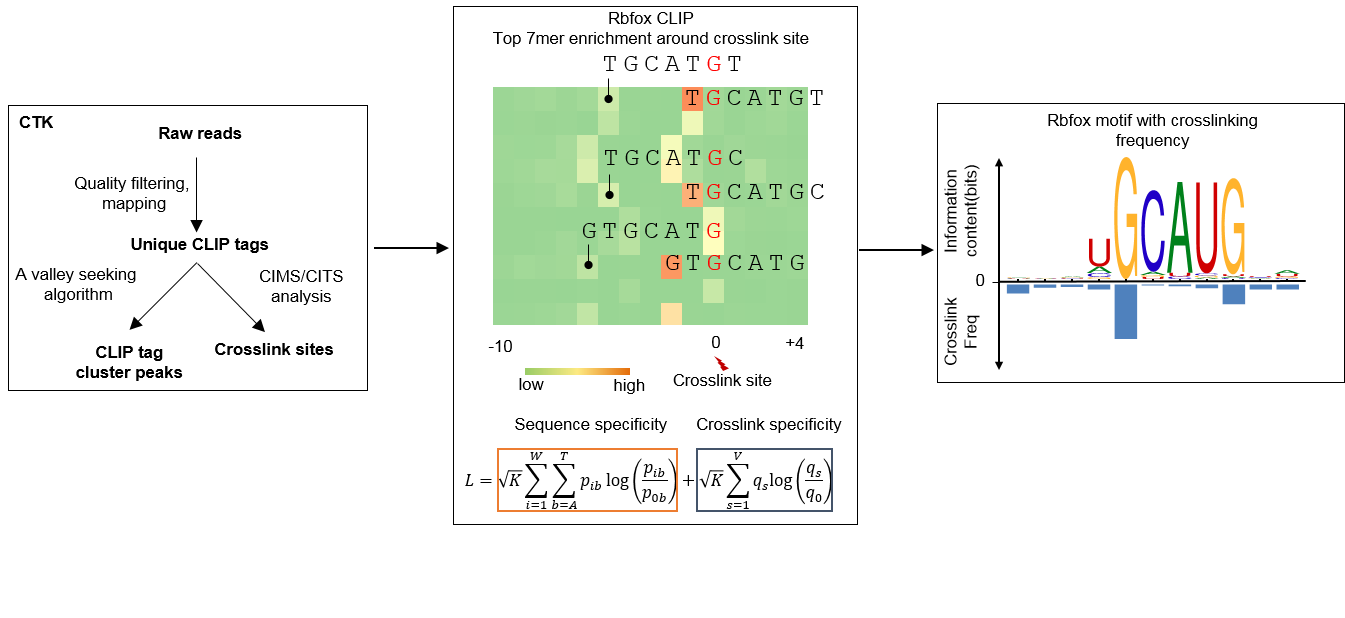
Framework to construct POSTAR2 database. (A) POSTAR2 covers six species... | Download Scientific Diagram

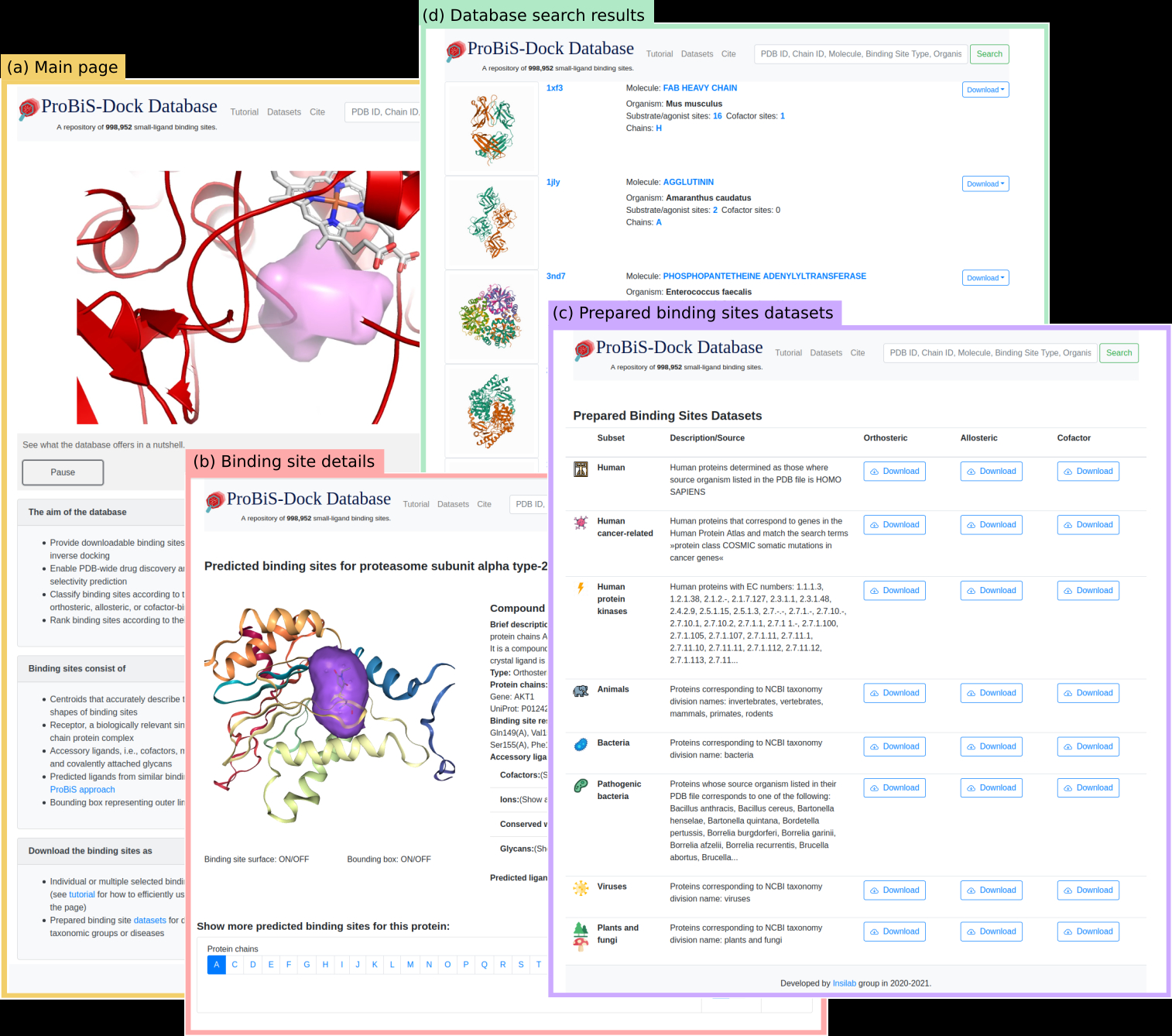
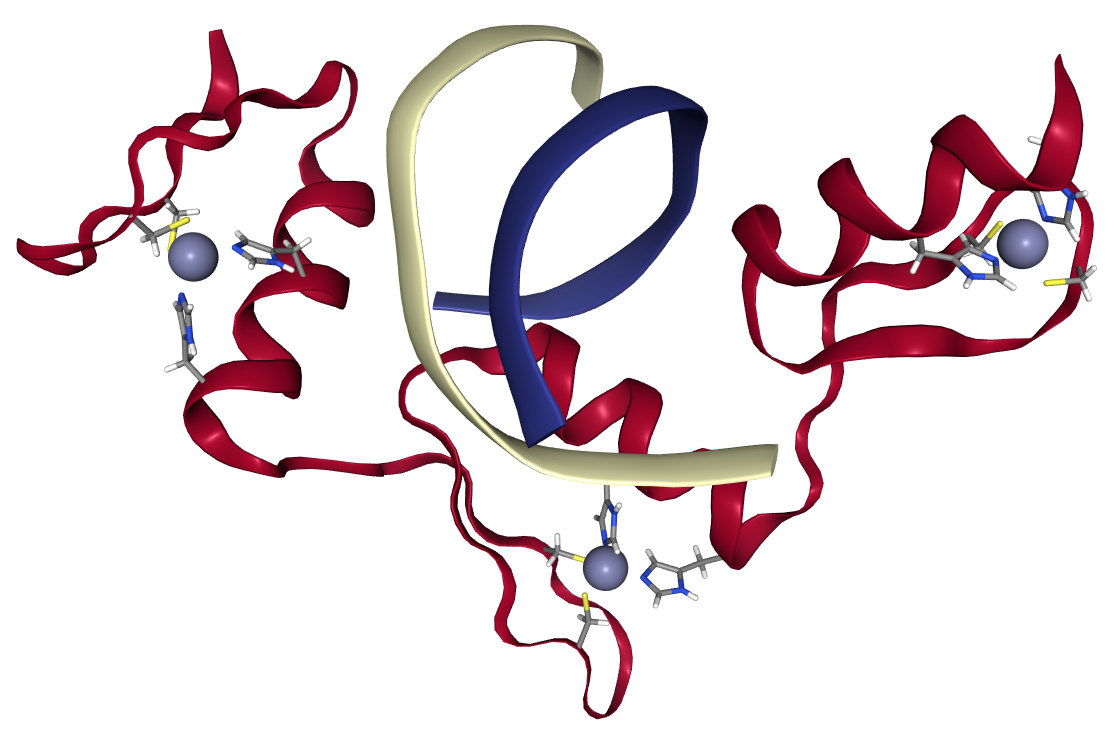
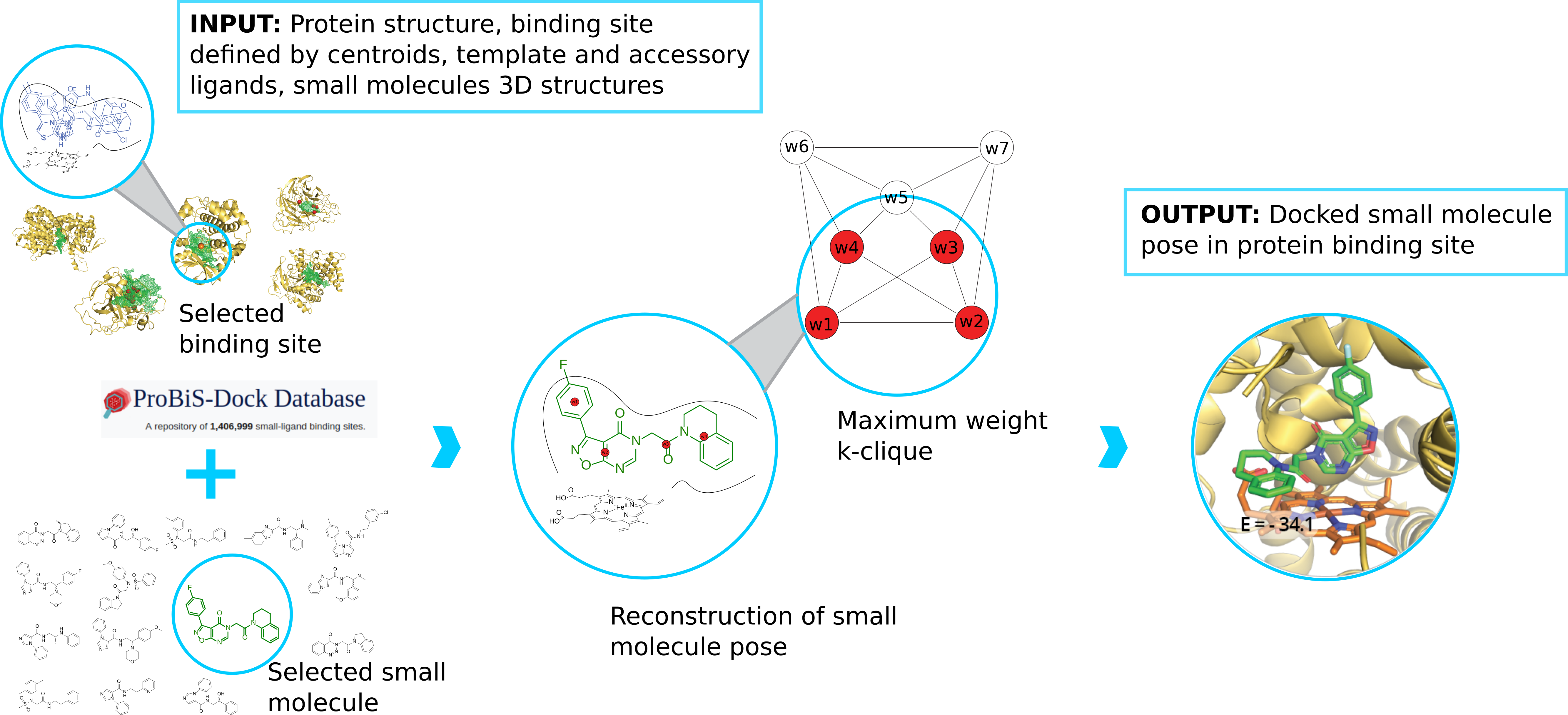
ProBiS-Dock Database: A Web Server and Interactive Web Repository of Small Ligand–Protein Binding Sites for Drug Design | Journal of Chemical Information and Modeling
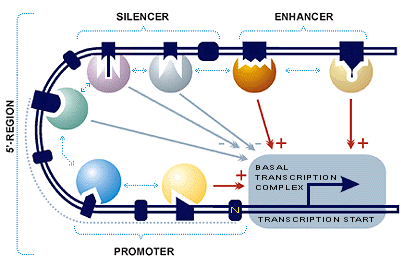
microPIR: An Integrated Database of MicroRNA Target Sites within Human Promoter Sequences | PLOS ONE

Biomolecules | Free Full-Text | CavitySpace: A Database of Potential Ligand Binding Sites in the Human Proteome
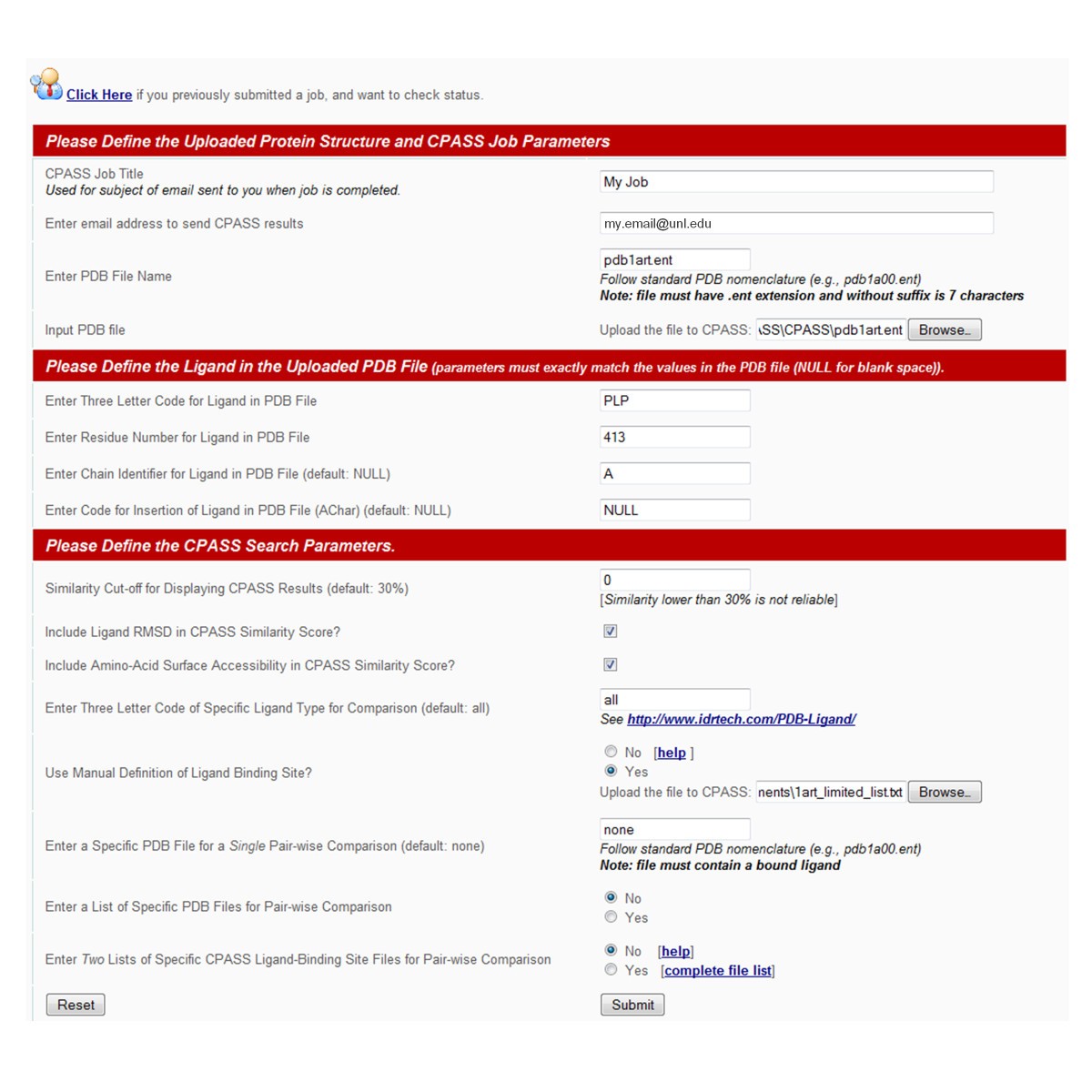
Searching the protein structure database for ligand-binding site similarities using CPASS v.2 | BMC Research Notes | Full Text