Alignment of DNA binding domain in LuxR regulators and prediction of... | Download Scientific Diagram
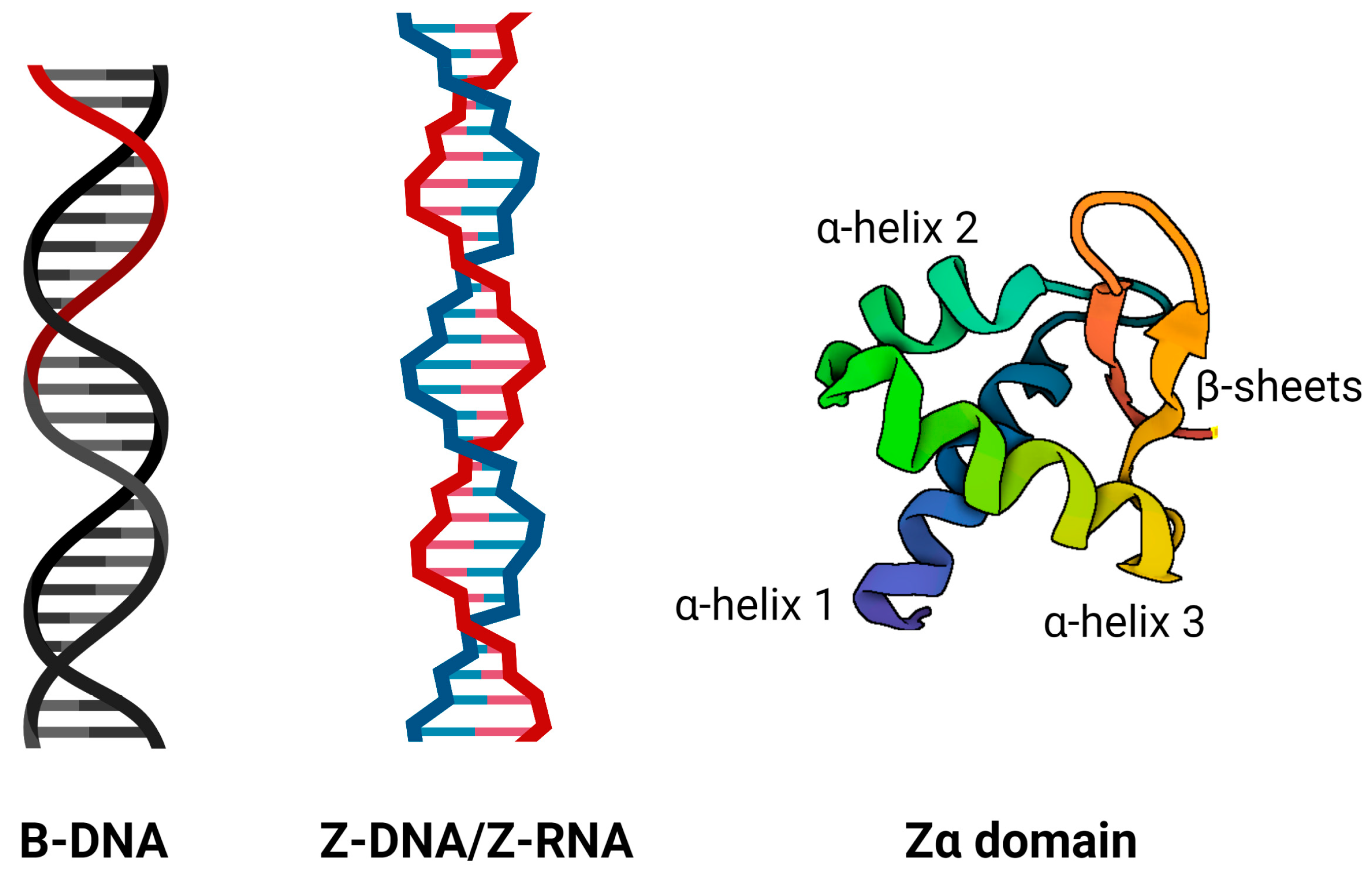
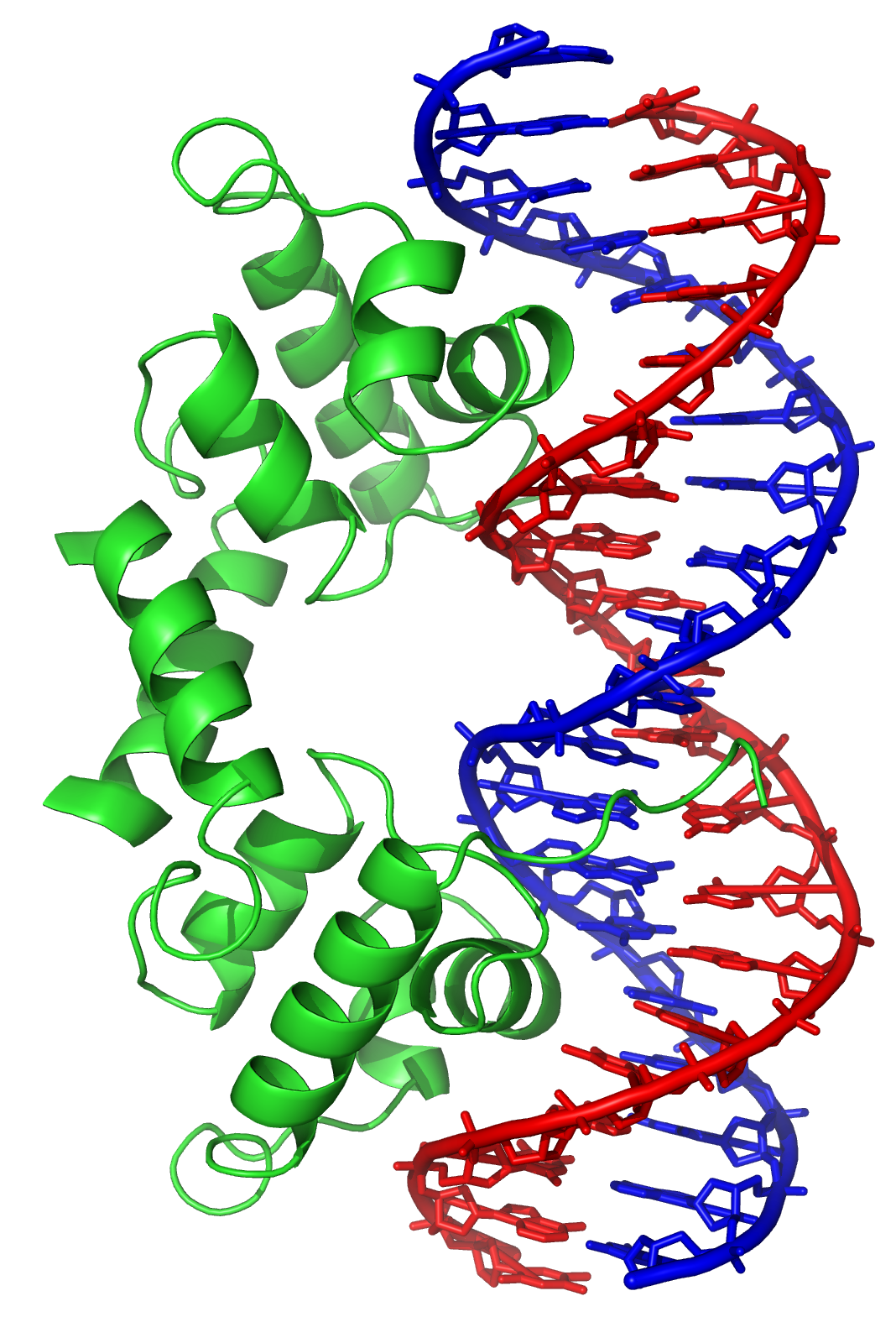
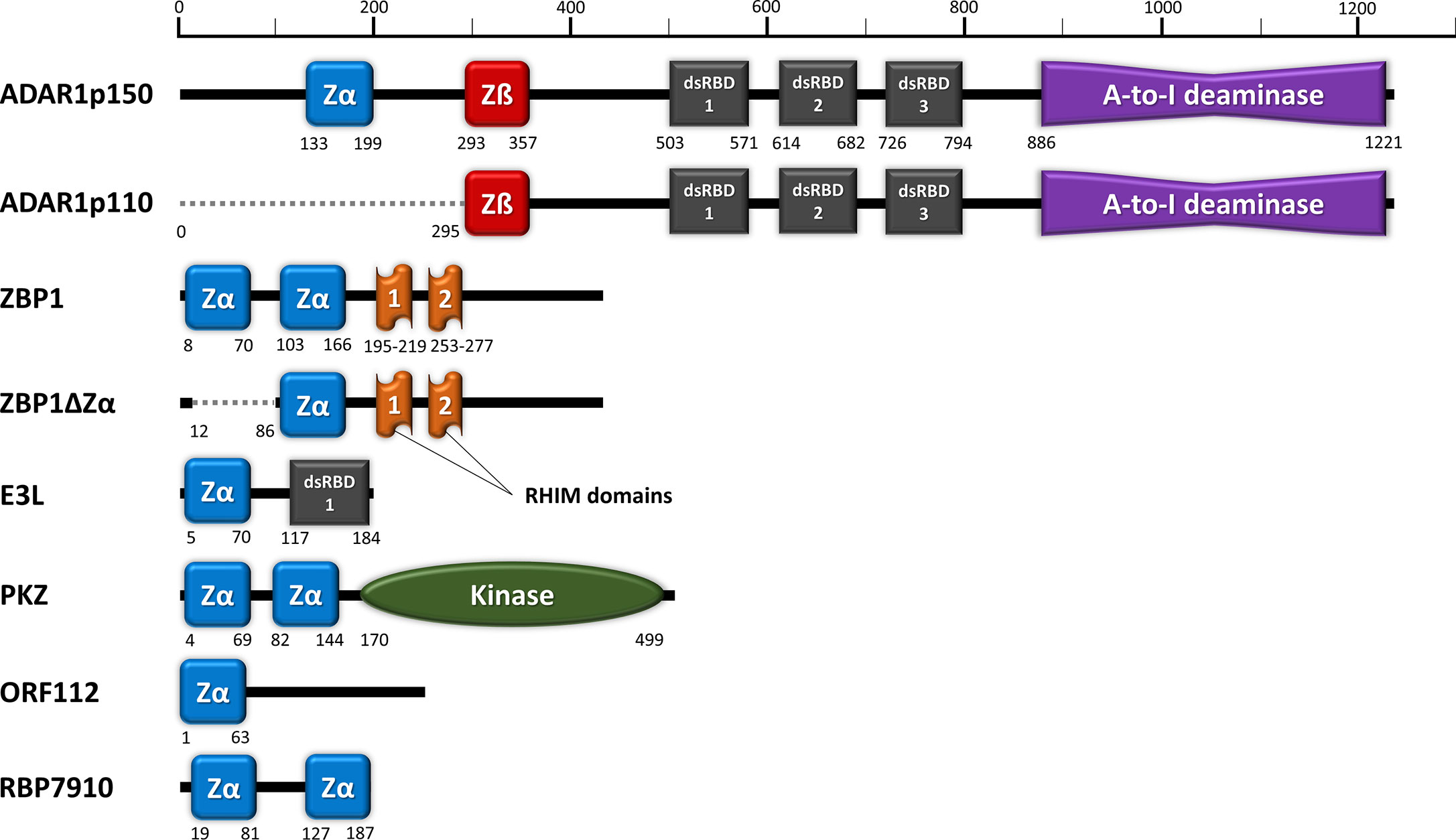
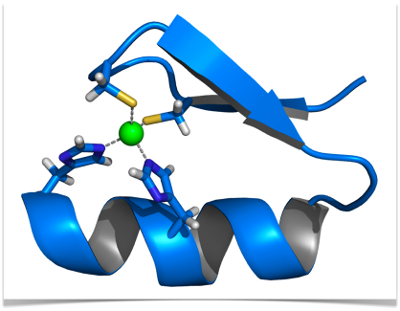
IJMS | Free Full-Text | Searching for New Z-DNA/Z-RNA Binding Proteins Based on Structural Similarity to Experimentally Validated Zα Domain

The Role of Charge Density Coupled DNA Bending in Transcription Factor Sequence Binding Specificity: A Generic Mechanism for Indirect Readout | Journal of the American Chemical Society

Using Differential Peak Prediction to Identify Motifs of Different DNA... | Download Scientific Diagram
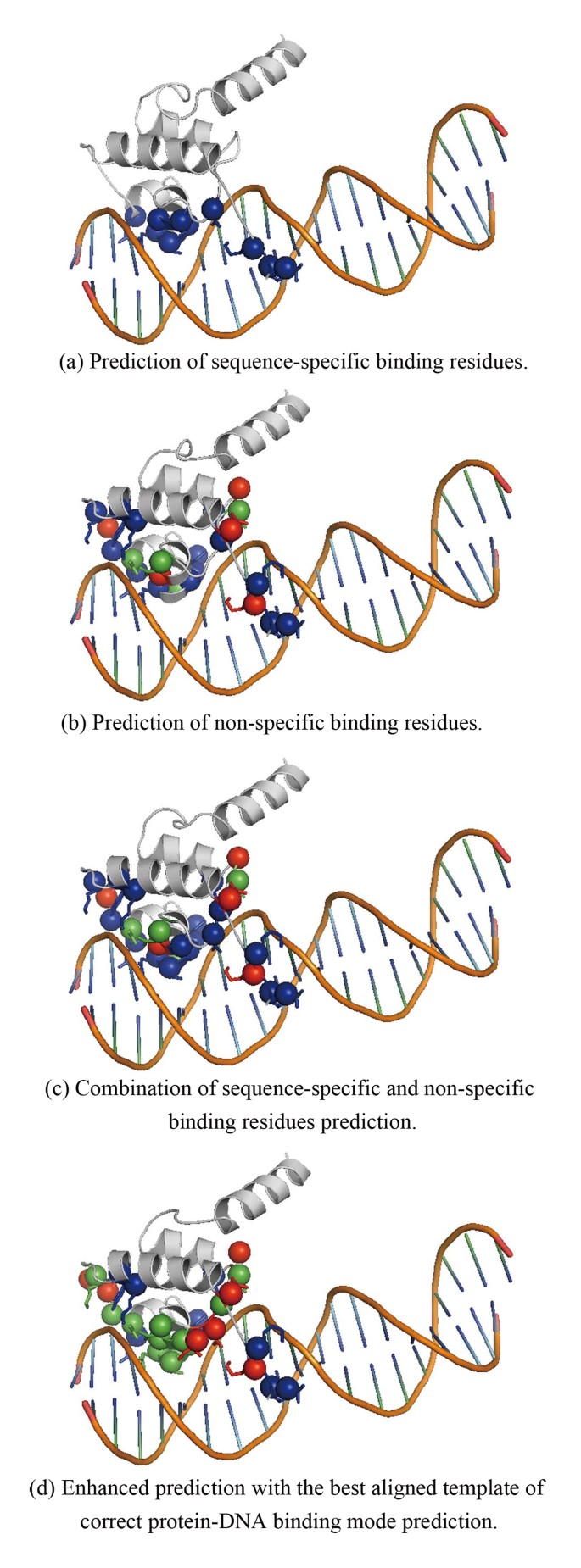
DNA-binding residues and binding mode prediction with binding-mechanism concerned models | BMC Genomics | Full Text

Predicting transcription factor binding sites using DNA shape features based on shared hybrid deep learning architecture: Molecular Therapy - Nucleic Acids
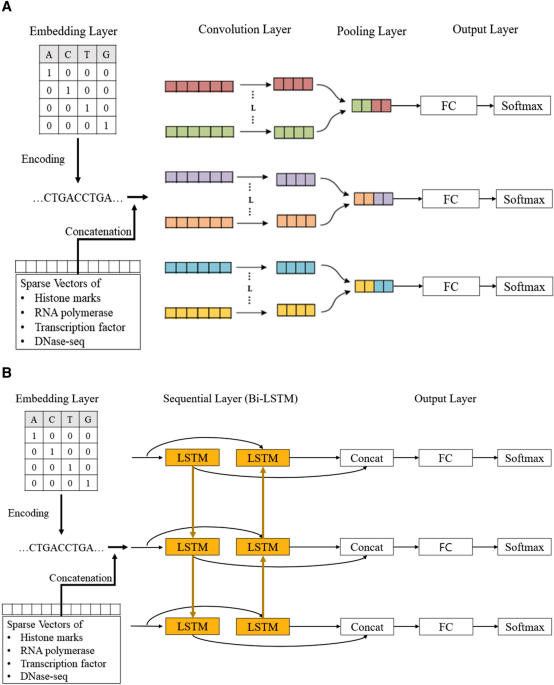
Deep learning approach for predicting functional Z-DNA regions using omics data | Scientific Reports
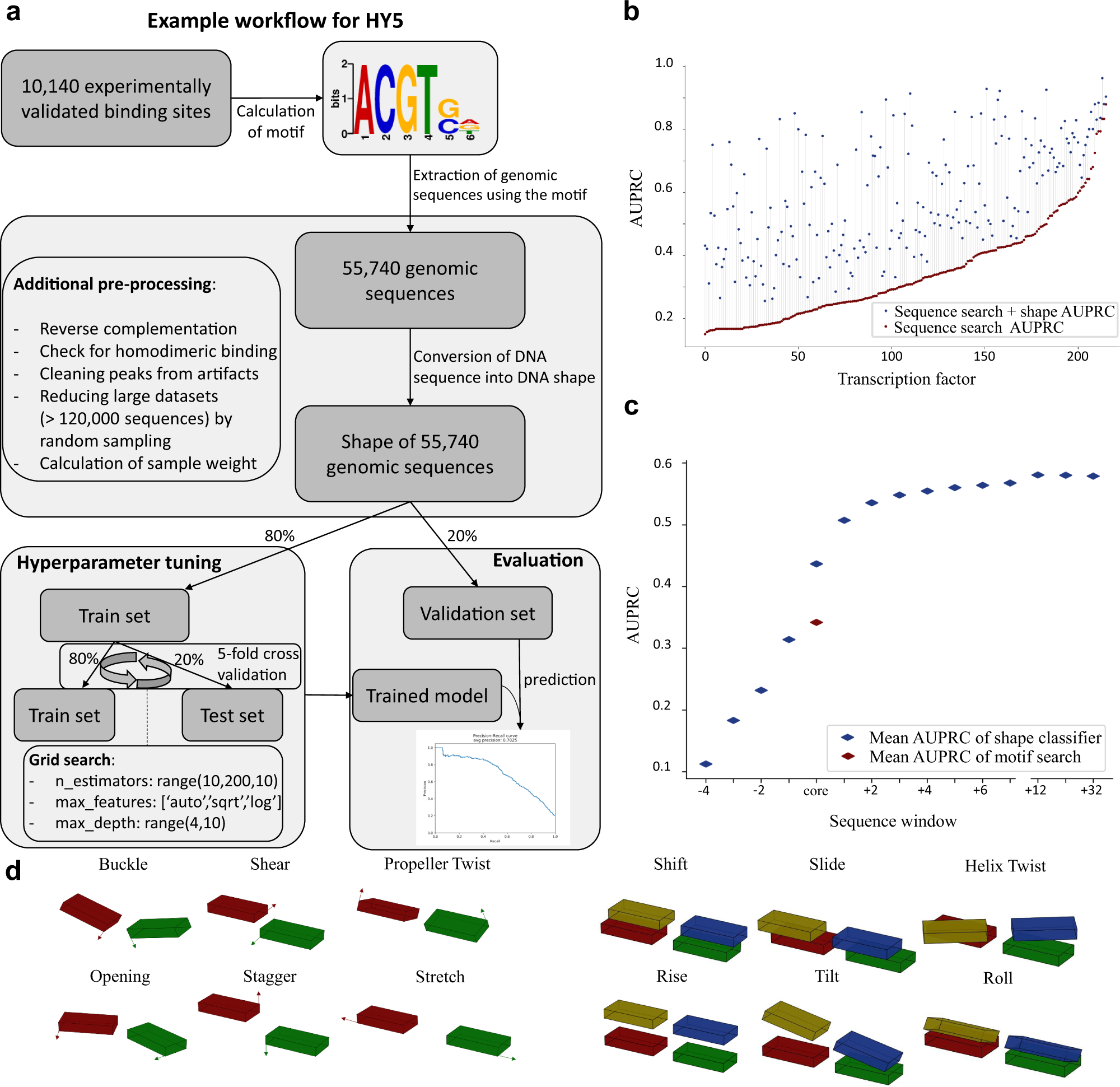
Local DNA shape is a general principle of transcription factor binding specificity in Arabidopsis thaliana | Nature Communications
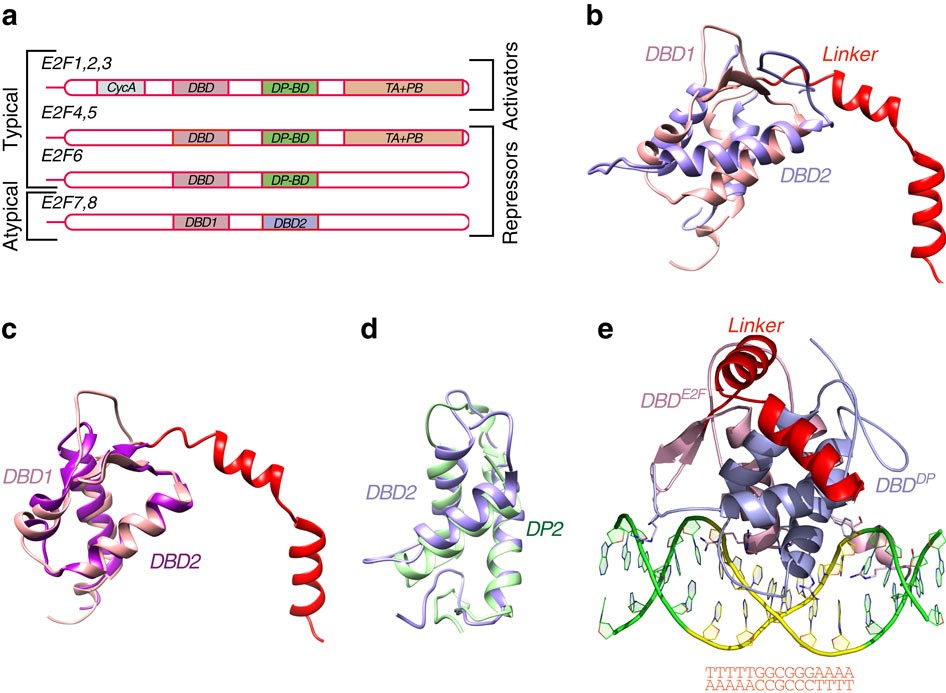
Structural insights into the DNA-binding specificity of E2F family transcription factors | Nature Communications
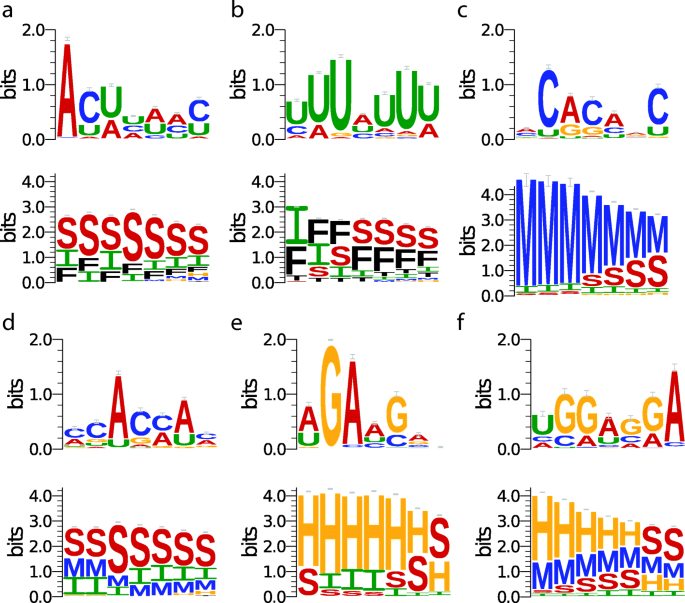
Prediction of RNA-protein sequence and structure binding preferences using deep convolutional and recurrent neural networks | BMC Genomics | Full Text

Examples of DNA-binding protein prediction on APO104. (A–D) In each... | Download Scientific Diagram
![Prediction of DNA binding proteins using local features and long-term dependencies with primary sequences based on deep learning [PeerJ] Prediction of DNA binding proteins using local features and long-term dependencies with primary sequences based on deep learning [PeerJ]](https://dfzljdn9uc3pi.cloudfront.net/2021/11262/1/fig-1-2x.jpg)
Prediction of DNA binding proteins using local features and long-term dependencies with primary sequences based on deep learning [PeerJ]

Protein–DNA/RNA Interactions: An Overview of Investigation Methods in the -Omics Era | Journal of Proteome Research

A compendium of DNA-binding specificities of transcription factors in Pseudomonas syringae | Nature Communications

The intervening domain is required for DNA-binding and functional identity of plant MADS transcription factors | Nature Communications