Affinity binding of antibodies to supermacroporous cryogel adsorbents with immobilized protein A for removal of anthrax toxin protective antigen - ScienceDirect

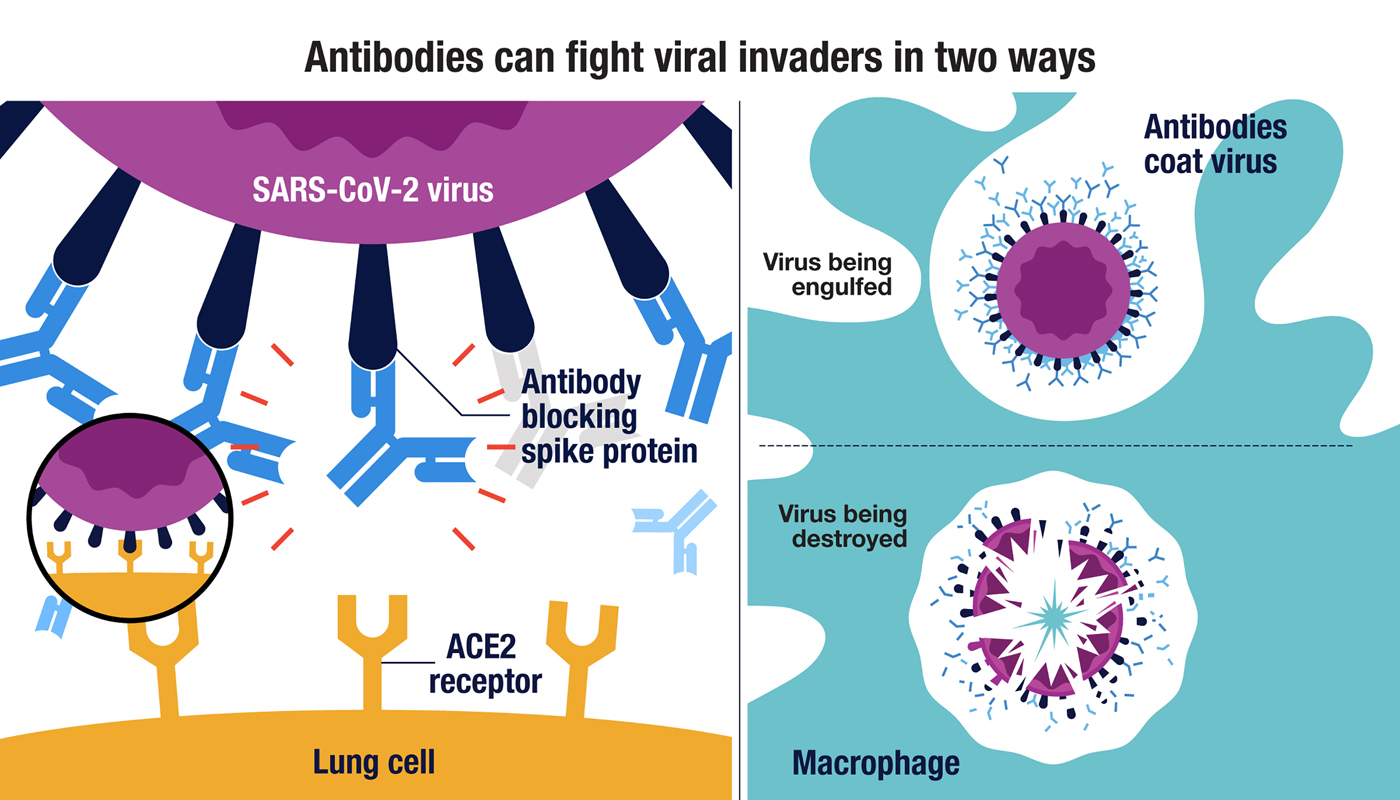
A human monoclonal antibody bivalently binding two different epitopes in streptococcal M protein mediates immune function | EMBO Molecular Medicine

Computational and artificial intelligence-based methods for antibody development: Trends in Pharmacological Sciences

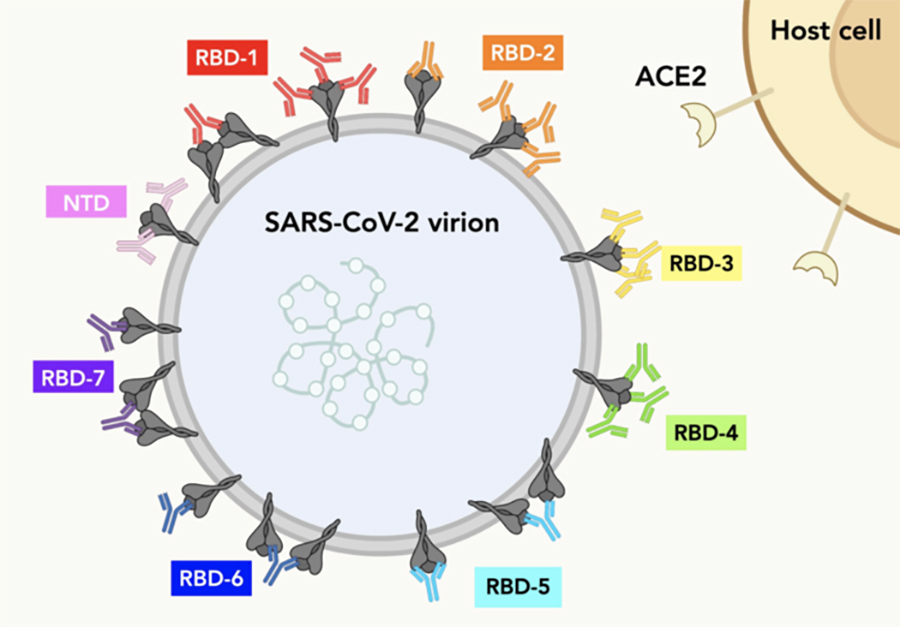
MHC-II constrains the natural neutralizing antibody response to the SARS-CoV-2 spike RBM in humans | bioRxiv
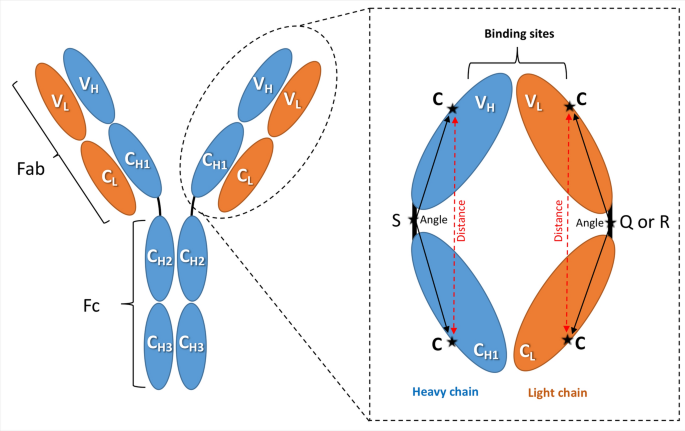
Antibody-protein binding and conformational changes: identifying allosteric signalling pathways to engineer a better effector response | Scientific Reports

Native Isolation of 3×HA‐Tagged Protein Complexes to Characterize Protein‐ Protein Interactions - Lim - 2021 - Current Protocols - Wiley Online Library